Journal Description
Viruses
Viruses
is a peer-reviewed, open access journal of virology, published monthly online by MDPI. The American Society for Virology (ASV), Spanish Society for Virology (SEV), Canadian Society for Virology (CSV), Italian Society for Virology (SIV-ISV), Australasian Virology Society (AVS) and others are affiliated with Viruses and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Virology) / CiteScore - Q1 (Infectious Diseases)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 13.8 days after submission; acceptance to publication is undertaken in 2.5 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Companion journals for Viruses include: COVID and Zoonotic Diseases.
Impact Factor:
4.7 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
The Interplay between KSHV Infection and DNA-Sensing Pathways
Viruses 2024, 16(5), 749; https://doi.org/10.3390/v16050749 - 8 May 2024
Abstract
During viral infection, the innate immune system utilizes a variety of specific intracellular sensors to detect virus-derived nucleic acids and activate a series of cellular signaling cascades that produce type Ⅰ IFNs and proinflammatory cytokines and chemokines. Kaposi’s sarcoma-associated herpesvirus (KSHV) is an
[...] Read more.
During viral infection, the innate immune system utilizes a variety of specific intracellular sensors to detect virus-derived nucleic acids and activate a series of cellular signaling cascades that produce type Ⅰ IFNs and proinflammatory cytokines and chemokines. Kaposi’s sarcoma-associated herpesvirus (KSHV) is an oncogenic double-stranded DNA virus that has been associated with a variety of human malignancies, including Kaposi’s sarcoma, primary effusion lymphoma, and multicentric Castleman disease. Infection with KSHV activates various DNA sensors, including cGAS, STING, IFI16, and DExD/H-box helicases. Activation of these DNA sensors induces the innate immune response to antagonize the virus. To counteract this, KSHV has developed countless strategies to evade or inhibit DNA sensing and facilitate its own infection. This review summarizes the major DNA-triggered sensing signaling pathways and details the current knowledge of DNA-sensing mechanisms involved in KSHV infection, as well as how KSHV evades antiviral signaling pathways to successfully establish latent infection and undergo lytic reactivation.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
Open AccessArticle
Phylogenetic Relationships and Evolution of the Genus Eganvirus (186-Type) Yersinia pestis Bacteriophages
by
Jin Guo, Youhong Zhong, Yiting Wang, Pan Liu, Haixiao Jin, Yumeng Wang, Liyuan Shi, Peng Wang and Wei Li
Viruses 2024, 16(5), 748; https://doi.org/10.3390/v16050748 - 8 May 2024
Abstract
Plague is an endemic infectious disease caused by Yersinia pestis. In this study, we isolated fourteen phages with similar sequence arrangements to phage 186; these phages exhibited different lytic abilities in Enterobacteriaceae strains. To illustrate the phylogenetic relationships and evolutionary relationships between
[...] Read more.
Plague is an endemic infectious disease caused by Yersinia pestis. In this study, we isolated fourteen phages with similar sequence arrangements to phage 186; these phages exhibited different lytic abilities in Enterobacteriaceae strains. To illustrate the phylogenetic relationships and evolutionary relationships between previously designated 186-type phages, we analysed the complete sequences and important genes of the phages, including whole-genome average nucleotide identity (ANI) and collinearity comparison, evolutionary analysis of four conserved structural genes (V, T, R, and Q genes), and analysis of the regulatory genes (cI, apl, and cII) and integrase gene (int). Phylogenetic analysis revealed that thirteen of the newly isolated phages belong to the genus Eganvirus and one belongs to the genus Felsduovirus in the family Peduoviridae, and these Eganvirus phages can be roughly clustered into three subgroups. The topological relationships exhibited by the whole-genome and structural genes seemed similar and stable, while the regulatory genes presented different topological relationships with the structural genes, and these results indicated that there was some homologous recombination in the regulatory genes. These newly isolated 186-type phages were mostly isolated from dogs, suggesting that the resistance of Canidae to Y. pestis infection may be related to the wide distribution of phages with lytic capability.
Full article
(This article belongs to the Special Issue Bacteriophage Diversity)
Open AccessReview
Models of Herpes Simplex Virus Latency
by
Paige N. Canova, Audra J. Charron and David A. Leib
Viruses 2024, 16(5), 747; https://doi.org/10.3390/v16050747 - 8 May 2024
Abstract
Our current understanding of HSV latency is based on a variety of clinical observations, and in vivo, ex vivo, and in vitro model systems, each with unique advantages and drawbacks. The criteria for authentically modeling HSV latency include the ability to easily
[...] Read more.
Our current understanding of HSV latency is based on a variety of clinical observations, and in vivo, ex vivo, and in vitro model systems, each with unique advantages and drawbacks. The criteria for authentically modeling HSV latency include the ability to easily manipulate host genetics and biological pathways, as well as mimicking the immune response and viral pathogenesis in human infections. Although realistically modeling HSV latency is necessary when choosing a model, the cost, time requirement, ethical constraints, and reagent availability are also equally important. Presently, there remains a pressing need for in vivo models that more closely recapitulate human HSV infection. While the current in vivo, ex vivo, and in vitro models used to study HSV latency have limitations, they provide further insights that add to our understanding of latency. In vivo models have shed light on natural infection routes and the interplay between the host immune response and the virus during latency, while in vitro models have been invaluable in elucidating molecular pathways involved in latency. Below, we review the relative advantages and disadvantages of current HSV models and highlight insights gained through each.
Full article
(This article belongs to the Special Issue Advances in HSV Research)
Open AccessArticle
A Screening Study Identified Decitabine as an Inhibitor of Equid Herpesvirus 4 That Enhances the Innate Antiviral Response
by
Camille Normand, Côme J. Thieulent, Christine Fortier, Gabrielle Sutton, Catherine Senamaud-Beaufort, Laurent Jourdren, Corinne Blugeon, Pierre-Olivier Vidalain, Stéphane Pronost and Erika S. Hue
Viruses 2024, 16(5), 746; https://doi.org/10.3390/v16050746 - 8 May 2024
Abstract
Equid herpesvirus 4 (EHV-4) is a common respiratory pathogen in horses. It sporadically induces abortion or neonatal death. Although its contribution in neurological disorders is not clearly demonstrated, there is a strong suspicion of its involvement. Despite preventive treatments using vaccines against EHV-1/EHV-4,
[...] Read more.
Equid herpesvirus 4 (EHV-4) is a common respiratory pathogen in horses. It sporadically induces abortion or neonatal death. Although its contribution in neurological disorders is not clearly demonstrated, there is a strong suspicion of its involvement. Despite preventive treatments using vaccines against EHV-1/EHV-4, the resurgence of alpha-EHV infection still constitutes an important threat to the horse industry. Yet very few studies have been conducted on the search for antiviral molecules against EHV-4. A screening of 42 antiviral compounds was performed in vitro on equine fibroblast cells infected with the EHV-4 405/76 reference strain (VR2230). The formation of cytopathic effects was monitored by real-time cell analysis (RTCA), and the viral load was quantified by quantitative PCR. Aciclovir, the most widely used antiviral against alpha-herpesviruses in vivo, does not appear to be effective against EHV-4 in vitro. Potential antiviral activities were confirmed for eight molecules (idoxuridine, vidarabine, pritelivir, cidofovir, valganciclovir, ganciclovir, aphidicolin, and decitabine). Decitabine demonstrates the highest efficacy against EHV-4 in vitro. Transcriptomic analysis revealed the up-regulation of various genes implicated in interferon (IFN) response, suggesting that decitabine triggers the immune antiviral pathway.
Full article
(This article belongs to the Special Issue Viral Cycle and Cell Host Interactions of Equine Viruses)
►▼
Show Figures

Figure 1
Open AccessReview
Roles Played by DOCK11, a Guanine Nucleotide Exchange Factor, in HBV Entry and Persistence in Hepatocytes
by
Ying-Yi Li, Kazuhisa Murai, Junyan Lyu and Masao Honda
Viruses 2024, 16(5), 745; https://doi.org/10.3390/v16050745 - 8 May 2024
Abstract
HBV infection is challenging to cure due to the persistence of viral covalently closed circular viral DNA (cccDNA). The dedicator of cytokinesis 11 (DOCK11) is recognized as a guanine nucleotide exchange factor (GEF) for CDC42 that has been reported to be required for
[...] Read more.
HBV infection is challenging to cure due to the persistence of viral covalently closed circular viral DNA (cccDNA). The dedicator of cytokinesis 11 (DOCK11) is recognized as a guanine nucleotide exchange factor (GEF) for CDC42 that has been reported to be required for HBV persistence. DOCK11 is expressed in both the cytoplasm and nucleus of human hepatocytes and is functionally associated with retrograde trafficking proteins Arf-GAP with GTPase domain, ankyrin repeat, and pleckstrin homology domain-containing protein 2 (AGAP2), and ADP-ribosylation factor 1 (ARF1), together with the HBV capsid, in the trans-Golgi network (TGN). This opens an alternative retrograde trafficking route for HBV from early endosomes (EEs) to the TGN and then to the endoplasmic reticulum (ER), thereby avoiding lysosomal degradation. DOCK11 also facilitates the association of cccDNA with H3K4me3 and RNA Pol II for activating cccDNA transcription. In addition, DOCK11 plays a crucial role in the host DNA repair system, being essential for cccDNA synthesis. This function can be inhibited by 10M-D42AN, a novel DOCK11-binding peptide, leading to the suppression of HBV replication both in vitro and in vivo. Treatment with a combination of 10M-D42AN and entecavir may represent a promising therapeutic strategy for patients with chronic hepatitis B (CHB). Consequently, DOCK11 may be seen as a potential candidate molecule in the development of molecularly targeted drugs against CHB.
Full article
(This article belongs to the Special Issue Unraveling the Pathogenesis of Persistent Virus Infection)
►▼
Show Figures
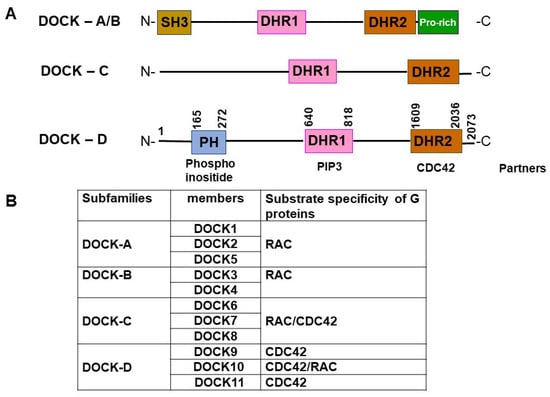
Figure 1
Open AccessArticle
Prevalence of Acute Hepatitis E Virus Infections in Swiss Blood Donors 2018–2020
by
Christoph Niederhauser, Peter Gowland, Nadja Widmer, Soraya Amar EL Dusouqui, Maja Mattle-Greminger, Jochen Gottschalk and Beat M. Frey
Viruses 2024, 16(5), 744; https://doi.org/10.3390/v16050744 - 8 May 2024
Abstract
Introduction: Hepatitis E virus (HEV) genotype 3 is the major cause of acute viral hepatitis in several European countries. It is acquired mainly by ingesting contaminated pork, but has also been reported to be transmitted through blood transfusion. Although most HEV infections, including
[...] Read more.
Introduction: Hepatitis E virus (HEV) genotype 3 is the major cause of acute viral hepatitis in several European countries. It is acquired mainly by ingesting contaminated pork, but has also been reported to be transmitted through blood transfusion. Although most HEV infections, including those via blood products, are usually self-limiting, they may become chronic in immunocompromised persons. It is thus essential to identify HEV-infected blood donations to prevent transmission to vulnerable recipients. Aims: Prior to the decision whether to introduce HEV RNA screening for all Swiss blood donations, a 2-year nationwide prevalence study was conducted. Methods: All blood donations were screened in pools of 12–24 samples at five regional blood donation services, and HEV RNA-positive pools were subsequently resolved to the individual donation index donation (X). The viral load, HEV IgG and IgM serology, and HEV genotype were determined. Follow-up investigations were conducted on future control donations (X + 1) and previous archived donations of the donor (X − 1) where available. Results: Between October 2018 and September 2020, 541,349 blood donations were screened and 125 confirmed positive donations were identified (prevalence 1:4331 donations). At the time of blood donation, the HEV RNA-positive individuals were symptom-free. The median viral load was 554 IU/mL (range: 2.01–2,500,000 IU/mL). Men (88; 70%) were more frequently infected than women (37; 30%), as compared with the sex distribution in the Swiss donor population (57% male/43% female, p < 0.01). Of the 106 genotyped cases (85%), all belonged to genotype 3. Two HEV sub-genotypes predominated; 3h3 (formerly 3s) and 3c. The remaining sub-genotypes are all known to circulate in Europe. Five 3ra genotypes were identified, this being a variant associated with rabbits. In total, 85 (68%) X donations were negative for HEV IgM and IgG. The remaining 40 (32%) were positive for HEV IgG and/or IgM, and consistent with an active infection. We found no markers of previous HEV in 87 of the 89 available and analyzed archive samples (X − 1). Two donors were HEV IgG-positive in the X − 1 donation suggesting insufficient immunity to prevent HEV reinfection. Time of collection of the 90 (72%) analyzed X + 1 donations varied between 2.9 and 101.9 weeks (median of 35 weeks) after X donation. As expected, none of those tested were positive for HEV RNA. Most donors (89; 99%) were positive for anti-HEV lgG/lgM (i.e., seroconversion). HEV lgM-positivity (23; 26%) indicates an often-long persistence of lgM antibodies post-HEV infection. Conclusion: The data collected during the first year of the study provided the basis for the decision to establish mandatory HEV RNA universal screening of all Swiss blood donations in minipools, a vital step in providing safer blood for all recipients, especially those who are immunosuppressed.
Full article
(This article belongs to the Special Issue Epidemiology and Diagnostics of Hepatitis Viruses)
►▼
Show Figures

Figure 1
Open AccessArticle
Phage–Bacterial Interaction Alters Phenotypes Associated with Virulence in Acinetobacter baumannii
by
Greater Kayode Oyejobi, Xiaoxu Zhang, Dongyan Xiong, Heng Xue, Mengjuan Shi, Hang Yang and Hongping Wei
Viruses 2024, 16(5), 743; https://doi.org/10.3390/v16050743 - 8 May 2024
Abstract
Bacteriophages exert strong selection on their bacterial hosts to evolve resistance. At the same time, the fitness costs on bacteria following phage resistance may change their virulence, which may affect the therapeutic outcomes of phage therapy. In this study, we set out to
[...] Read more.
Bacteriophages exert strong selection on their bacterial hosts to evolve resistance. At the same time, the fitness costs on bacteria following phage resistance may change their virulence, which may affect the therapeutic outcomes of phage therapy. In this study, we set out to assess the costs of phage resistance on the in vitro virulence of priority 1 nosocomial pathogenic bacterium, Acinetobacter baumannii. By subjecting phage-resistant variant Ev5-WHG of A. baumannii WHG40004 to several in vitro virulence profiles, we found that its resistance to phage is associated with reduced fitness in host microenvironments. Also, the mutant exhibited impaired adhesion and invasion to mammalian cells, as well as increased susceptibility to macrophage phagocytosis. Furthermore, the whole-genome sequencing of the mutant revealed that there exist multiple mutations which may play a role in phage resistance and altered virulence. Altogether, this study demonstrates that resistance to phage can significantly alter phenotypes associated with virulence in Acinetobacter baumannii.
Full article
(This article belongs to the Special Issue Phage-Bacteria Interplay in Health and Disease, Second Edition)
►▼
Show Figures
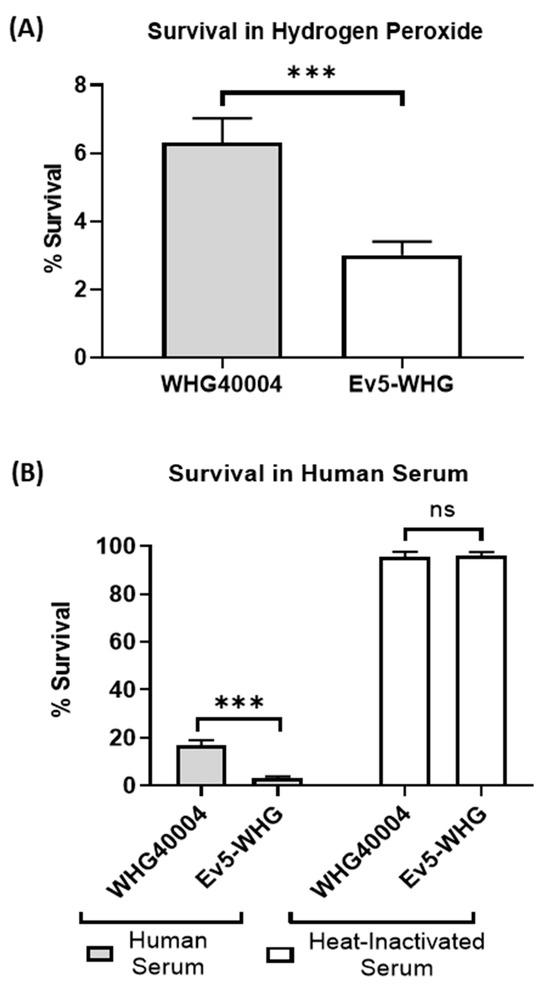
Figure 1
Open AccessCorrection
Correction: Wang et al. Nicotinic Acetylcholine Receptor Alpha6 Contributes to Antiviral Immunity via IMD Pathway in Drosophila melanogaster. Viruses 2024, 16, 562
by
Zhiying Wang, Xiaoju Lin, Wangpeng Shi and Chuan Cao
Viruses 2024, 16(5), 742; https://doi.org/10.3390/v16050742 - 8 May 2024
Abstract
In the original publication [...]
Full article
(This article belongs to the Special Issue Molecular Virus-Insect Interactions)
Open AccessArticle
Cytokine Response of Natural Killer Cells to Hepatitis B Virus Infection Depends on Monocyte Co-Stimulation
by
Paul Kupke, Johanna Brucker, Jochen M. Wettengel, Ulrike Protzer, Jürgen J. Wenzel, Hans J. Schlitt, Edward K. Geissler and Jens M. Werner
Viruses 2024, 16(5), 741; https://doi.org/10.3390/v16050741 - 8 May 2024
Abstract
Hepatitis B virus (HBV) is a major driver of chronic hepatic inflammation, which regularly leads to liver cirrhosis or hepatocellular carcinoma. Immediate innate immune cell response is crucial for the rapid clearance of the infection. Here, natural killer (NK) cells play a pivotal
[...] Read more.
Hepatitis B virus (HBV) is a major driver of chronic hepatic inflammation, which regularly leads to liver cirrhosis or hepatocellular carcinoma. Immediate innate immune cell response is crucial for the rapid clearance of the infection. Here, natural killer (NK) cells play a pivotal role in direct cytotoxicity and the secretion of antiviral cytokines as well as regulatory function. The aim of this study was to further elucidate NK cell responses triggered by an HBV infection. Therefore, we optimized HBV in vitro models that reliably stimulate NK cells using hepatocyte-like HepG2 cells expressing the Na+-taurocholate co-transporting polypeptide (NTCP) and HepaRG cells. Immune cells were acquired from healthy platelet donors. Initially, HepG2-NTCP cells demonstrated higher viral replication compared to HepaRG cells. Co-cultures with immune cells revealed increased production of interferon-γ and tumor necrosis factor-α by NK cells, which was no longer evident in isolated NK cells. Likewise, the depletion of monocytes and spatial separation from target cells led to the absence of the antiviral cytokine production of NK cells. Eventually, the combined co-culture of isolated NK cells and monocytes led to a sufficient cytokine response of NK cells, which was also apparent when communication between the two immune cell subpopulations was restricted to soluble factors. In summary, our study demonstrates antiviral cytokine production by NK cells in response to HBV+ HepG2-NTCP cells, which is dependent on monocyte bystander activation.
Full article
(This article belongs to the Special Issue Natural Killer Cell in Viral Infection)
►▼
Show Figures

Figure 1
Open AccessReview
Hepatocyte Intrinsic Innate Antiviral Immunity against Hepatitis Delta Virus Infection: The Voices of Bona Fide Human Hepatocytes
by
Yein Woo, Muyuan Ma, Masashi Okawa and Takeshi Saito
Viruses 2024, 16(5), 740; https://doi.org/10.3390/v16050740 - 8 May 2024
Abstract
The pathogenesis of viral infection is attributed to two folds: intrinsic cell death pathway activation due to the viral cytopathic effect, and immune-mediated extrinsic cellular injuries. The immune system, encompassing both innate and adaptive immunity, therefore acts as a double-edged sword in viral
[...] Read more.
The pathogenesis of viral infection is attributed to two folds: intrinsic cell death pathway activation due to the viral cytopathic effect, and immune-mediated extrinsic cellular injuries. The immune system, encompassing both innate and adaptive immunity, therefore acts as a double-edged sword in viral infection. Insufficient potency permits pathogens to establish lifelong persistent infection and its consequences, while excessive activation leads to organ damage beyond its mission to control viral pathogens. The innate immune response serves as the front line of defense against viral infection, which is triggered through the recognition of viral products, referred to as pathogen-associated molecular patterns (PAMPs), by host cell pattern recognition receptors (PRRs). The PRRs–PAMPs interaction results in the induction of interferon-stimulated genes (ISGs) in infected cells, as well as the secretion of interferons (IFNs), to establish a tissue-wide antiviral state in an autocrine and paracrine manner. Cumulative evidence suggests significant variability in the expression patterns of PRRs, the induction potency of ISGs and IFNs, and the IFN response across different cell types and species. Hence, in our understanding of viral hepatitis pathogenesis, insights gained through hepatoma cell lines or murine-based experimental systems are uncertain in precisely recapitulating the innate antiviral response of genuine human hepatocytes. Accordingly, this review article aims to extract and summarize evidence made possible with bona fide human hepatocytes-based study tools, along with their clinical relevance and implications, as well as to identify the remaining gaps in knowledge for future investigations.
Full article
(This article belongs to the Special Issue Life Cycle of Hepatitis D Virus (HDV) and HDV-Like Agents)
►▼
Show Figures

Figure 1
Open AccessArticle
Whole Genome Sequence-Based Analysis of Bovine Gammaherpesvirus 4 Isolated from Bovine Abortions
by
Florencia Romeo, Maximiliano Joaquín Spetter, Susana Beatriz Pereyra, Pedro Edgardo Morán, Erika Analía González Altamiranda, Enrique Leopoldo Louge Uriarte, Anselmo Carlos Odeón, Sandra Elizabeth Pérez and Andrea Elizabeth Verna
Viruses 2024, 16(5), 739; https://doi.org/10.3390/v16050739 - 8 May 2024
Abstract
Bovine gammaherpesvirus 4 (BoGHV4) is a member of the Gammaherspivirinae subfamily, Rhadinovirus genus. Its natural host is the bovine, and it is prevalent among the global cattle population. Although the complete genome of BoGHV4 has been successfully sequenced, the functions of most of
[...] Read more.
Bovine gammaherpesvirus 4 (BoGHV4) is a member of the Gammaherspivirinae subfamily, Rhadinovirus genus. Its natural host is the bovine, and it is prevalent among the global cattle population. Although the complete genome of BoGHV4 has been successfully sequenced, the functions of most of its genes remain unknown. Currently, only six strains of BoGHV4, all belonging to Genotype 1, have been sequenced. This is the first report of the nearly complete genome of Argentinean BoGHV4 strains isolated from clinical cases of abortion, representing the first BoGHV4 Genotype 2 and 3 genomes described in the literature. Both Argentinean isolates presented the highest nt p-distance values, indicating a greater level of divergence. Overall, the considerable diversity observed in the complete genomes and open reading frames underscores the distinctiveness of both Argentinean isolates compared to the existing BoGHV4 genomes. These findings support previous studies that categorized the Argentinean BoGHV4 strains 07-435 and 10-154 as Genotypes 3 and 2, respectively. The inclusion of these sequences represents a significant expansion to the currently limited pool of BoGHV4 genomes while providing an important basis to increase the knowledge of local isolates.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures

Figure 1
Open AccessArticle
PSPC1 Binds to HCV IRES and Prevents Ribosomal Protein S5 Binding, Inhibiting Viral RNA Translation
by
Sachin Kumar Tripathi, Ashish Aneja, Teji Borgaonkar and Saumitra Das
Viruses 2024, 16(5), 738; https://doi.org/10.3390/v16050738 - 7 May 2024
Abstract
Hepatitis C virus (HCV) infects the human liver, and its chronic infection is one of the major causes of Hepatocellular carcinoma. Translation of HCV RNA is mediated by an Internal Ribosome Entry Site (IRES) element located in the 5’UTR of viral RNA. Several
[...] Read more.
Hepatitis C virus (HCV) infects the human liver, and its chronic infection is one of the major causes of Hepatocellular carcinoma. Translation of HCV RNA is mediated by an Internal Ribosome Entry Site (IRES) element located in the 5’UTR of viral RNA. Several RNA Binding proteins of the host interact with the HCV IRES and modulate its function. Here, we demonstrate that PSPC1 (Paraspeckle Component 1), an essential paraspeckle component, upon HCV infection is relocalized and interacts with HCV IRES to prevent viral RNA translation. Competition UV-crosslinking experiments showed that PSPC1 interacts explicitly with the SLIV region of the HCV IRES, which is known to play a vital role in ribosomal loading to the HCV IRES via interaction with Ribosomal protein S5 (RPS5). Partial silencing of PSPC1 increased viral RNA translation and, consequently, HCV replication, suggesting a negative regulation by PSPC1. Interestingly, the silencing of PSPC1 protein leads to an increased interaction of RPS5 at the SLIV region, leading to an overall increase in the viral RNA in polysomes. Overall, our results showed how the host counters viral infection by relocalizing nuclear protein to the cytoplasm as a survival strategy.
Full article
(This article belongs to the Special Issue Functional and Structural Features of Viral RNA Elements)
►▼
Show Figures
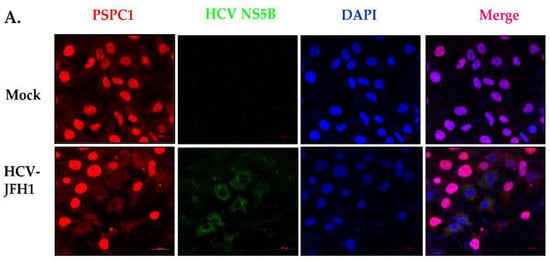
Figure 1
Open AccessArticle
Galectin-3-ITGB1 Signaling Mediates Interleukin 10 Production of Hepatic Conventional Natural Killer Cells in Chronic Hepatitis Virus B Transgenic Mice and Correlates with Hepatocellular Carcinoma Progression in Patients
by
Yongyan Chen, Wendi Zhang, Min Cheng, Xiaolei Hao, Haiming Wei, Rui Sun and Zhigang Tian
Viruses 2024, 16(5), 737; https://doi.org/10.3390/v16050737 - 7 May 2024
Abstract
Background and Aims: The outcomes of HBV infections are related to complex immune imbalances; however, the precise mechanisms by which HBV induces immune dysfunction are not well understood. Methods: HBV transgenic (HBs-Tg) mice were used to investigate intrahepatic NK cells in two distinct
[...] Read more.
Background and Aims: The outcomes of HBV infections are related to complex immune imbalances; however, the precise mechanisms by which HBV induces immune dysfunction are not well understood. Methods: HBV transgenic (HBs-Tg) mice were used to investigate intrahepatic NK cells in two distinct subsets: conventional NK (cNK) and liver-resident NK (LrNK) cells during a chronic HBV infection. Results: The cNK cells, but not the LrNK cells, were primarily responsible for the increase in the number of bulk NK cells in the livers of ageing HBs-Tg mice. The hepatic cNK cells showed a stronger ability to produce IL-10, coupled with a higher expression of CD69, TIGIT and PD-L1, and lower NKG2D expression in ageing HBs-Tg mice. A lower mitochondrial mass and membrane potential, and less polarized localization were observed in the hepatic cNK cells compared with the splenic cNK cells in the HBs-Tg mice. The enhanced galectin-3 (Gal-3) secreted from HBsAg+ hepatocytes accounted for the IL-10 production of hepatic cNK cells via ITGB1 signaling. For humans, LGALS3 and ITGB1 expression is positively correlated with IL-10 expression, and negatively correlated with the poor clinical progression of HCC. Conclusions: Gal-3-ITGB1 signaling shapes hepatic cNK cells but not LrNK cells during a chronic HBV infection, which may correlate with HCC progression.
Full article
(This article belongs to the Special Issue Natural Killer Cell in Viral Infection)
►▼
Show Figures

Figure 1
Open AccessArticle
Partial Alleviation of Homologous Superinfection Exclusion of SeMNPV Latently Infected Cells by G1 Phase Infection and G2/M Phase Arrest
by
Qi-Ming Fu, Zheng Fang, Lou Ren, Qing-Shan Wu, Jun-Bo Zhang, Qiu-Ping Liu, Lei-Tao Tan and Qing-Bei Weng
Viruses 2024, 16(5), 736; https://doi.org/10.3390/v16050736 - 6 May 2024
Abstract
Viral infection can regulate the cell cycle, thereby promoting viral replication. Hijacking and altering the cell cycle are important for the virus to establish and maintain a latent infection. Previously, Spodoptera exigua multiple nucleopolyhedrovirus (SeMNPV)-latently infected P8-Se301-C1 cells, which grew more slowly than
[...] Read more.
Viral infection can regulate the cell cycle, thereby promoting viral replication. Hijacking and altering the cell cycle are important for the virus to establish and maintain a latent infection. Previously, Spodoptera exigua multiple nucleopolyhedrovirus (SeMNPV)-latently infected P8-Se301-C1 cells, which grew more slowly than Se301 cells and interfered with homologous SeMNNPV superinfection, were established. However, the effects of latent and superinfection with baculoviruses on cell cycle progression remain unknown. In this study, the cell cycle profiles of P8-Se301-C1 cells and SeMNPV or Autographa californica multiple nucleopolyhedrovirus (AcMNPV)-infected P8-Se301-C1 cells were characterized by flow cytometry. The results showed that replication-related genes MCM4, PCNA, and BAF were down-regulated (p < 0.05) in P8-Se301-C1 cells, and the S phase of P8-Se301-C1 cells was longer than that of Se301 cells. P8-Se301-C1 cells infected with SeMNPV did not arrest in the G2/M phase or affect the expression of Cyclin B and cyclin-dependent kinase 1 (CDK1). Furthermore, when P8-Se301-C1 cells were infected with SeMNPV after synchronized treatment with hydroxyurea and nocodazole, light microscopy and qRT-PCR analysis showed that, compared with unsynchronized cells and S and G2/M phase cells, SeMNPV-infected P8-Se301-C1 cells in G1 phase induced G2/M phase arrest, and the amount of virus adsorption and intracellular viral DNA replication were significantly increased (p < 0.05). In addition, budded virus (BV) production and occlusion body (OB)-containing cells were both increased at 120 h post-infection (p < 0.05). The expression of Cyclin B and CDK1 was significantly down-regulated at 48 h post-infection (p < 0.05). Finally, the arrest of SeMNPV-infected G1 phase cells in the G2/M phase increased BV production (p < 0.05) and the number of OB-containing cells. In conclusion, G1 phase infection and G2/M arrest are favorable to SeMNPV proliferation in P8-Se301-C1 cells, thereby alleviating the homologous superinfection exclusion. The results contribute to a better understanding of the relationship between baculoviruses and insect cell cycle progression and regulation.
Full article
(This article belongs to the Special Issue Molecular Virus-Insect Interactions)
Open AccessArticle
Molecular Surveillance, Prevalence, and Distribution of Cacao Infecting Badnavirus Species in Côte d’Ivoire and Ghana
by
George A. Ameyaw, Koffié Kouakou, Mohammed Javed Iqbal, Luc Belé, Valentin L. F. Wolf, Cory V. Keith, Bolou A. Bolou Bi, Christophe Kouamé, Donald Livingstone, Owusu Domfeh, Ebenezer A. Gyamera, Jean-Philippe Marelli and Judith K. Brown
Viruses 2024, 16(5), 735; https://doi.org/10.3390/v16050735 - 6 May 2024
Abstract
The cacao swollen shoot disease (CSSD) caused by a complex of badnavirus species presents a major challenge for cacao production in West Africa, especially Ghana and Côte d’Ivoire. In this study, CSSD species detection efficiency, diversity, and geographic distribution patterns in cacao plantations
[...] Read more.
The cacao swollen shoot disease (CSSD) caused by a complex of badnavirus species presents a major challenge for cacao production in West Africa, especially Ghana and Côte d’Ivoire. In this study, CSSD species detection efficiency, diversity, and geographic distribution patterns in cacao plantations in Ghana and Côte d’Ivoire were investigated through field surveillance, PCR detection assays, sequencing of positive amplicons, and phylogeographic clustering. Cumulatively, the detection efficiency of the tested CSSD primer sets that were targeting the movement protein domain of the virus ranged from 0.15% (CSSD-3 primer) to 66.91% (CSSD-1 primer) on all the symptomatic cacao leaf samples assessed. The identified CSSD species differed phylogenetically and overlapped in distribution, with the cacao swollen shoot Togo B virus (CSSTBV) (n = 588 sequences) being the most prevalent and widely distributed compared to the other CSSD species that were encountered in both countries. Geographically, the cacao swollen shoot CE virus (CSSCEV) species (n = 124 sequences) that was identified was largely restricted to the bordering regions of Ghana and Côte d’Ivoire. These results provide updated knowledge of the geographic distribution of the key CSSD species and their diagnostic efficiency and, thus, provide guidance in identifying locations for structured testing of cacao germplasm and optimal diagnostics for the predominant CSSD species in Ghana and Côte d’Ivoire.
Full article
(This article belongs to the Special Issue Plant Viruses and Their Vectors: Epidemiology and Control)
►▼
Show Figures
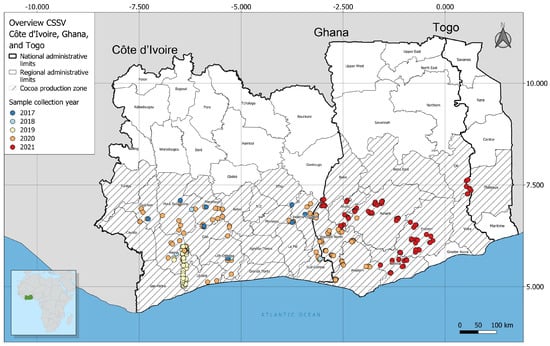
Figure 1
Open AccessReview
The Antiviral Activity of Interferon-Induced Transmembrane Proteins and Virus Evasion Strategies
by
Jingjing Wang, Yuhang Luo, Harshita Katiyar, Chen Liang and Qian Liu
Viruses 2024, 16(5), 734; https://doi.org/10.3390/v16050734 - 6 May 2024
Abstract
Interferons (IFNs) are antiviral cytokines that defend against viral infections by inducing the expression of interferon-stimulated genes (ISGs). Interferon-inducible transmembrane proteins (IFITMs) 1, 2, and 3 are crucial ISG products and members of the CD225 protein family. Compelling evidence shows that IFITMs restrict
[...] Read more.
Interferons (IFNs) are antiviral cytokines that defend against viral infections by inducing the expression of interferon-stimulated genes (ISGs). Interferon-inducible transmembrane proteins (IFITMs) 1, 2, and 3 are crucial ISG products and members of the CD225 protein family. Compelling evidence shows that IFITMs restrict the infection of many unrelated viruses by inhibiting the virus–cell membrane fusion at the virus entry step via the modulation of lipid composition and membrane properties. Meanwhile, viruses can evade IFITMs’ restrictions by either directly interacting with IFITMs via viral glycoproteins or by altering the native entry pathway. At the same time, cumulative evidence suggests context-dependent and multifaceted roles of IFITMs in modulating virus infections and cell signaling. Here, we review the diverse antiviral mechanisms of IFITMs, the viral antagonizing strategies, and the regulation of IFITM activity in host cells. The mechanisms behind the antiviral activity of IFITMs could aid the development of broad-spectrum antivirals and enhance preparedness for future pandemics.
Full article
(This article belongs to the Special Issue Interferon-Induced Transmembrane Proteins at the Intersection of Virus Infection and Immunity)
►▼
Show Figures
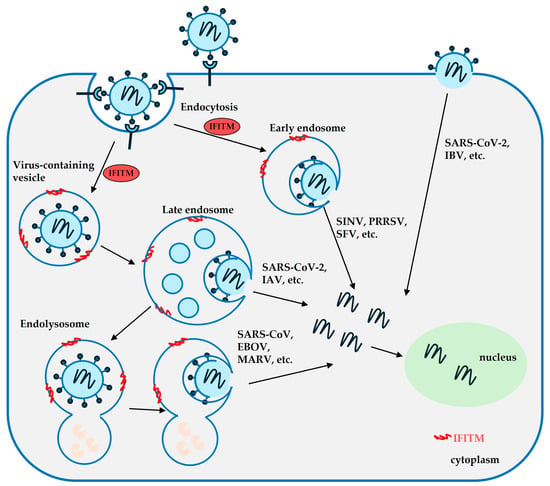
Figure 1
Open AccessEditorial
Pediatric Respiratory Viral Infection
by
Stacy L. S. Yam, Joan Marie Javillo Baguio and Renee W. Y. Chan
Viruses 2024, 16(5), 733; https://doi.org/10.3390/v16050733 - 6 May 2024
Abstract
Reflecting on this Special Issue dedicated to pediatric respiratory viruses, it is evident that the shadow cast by the global SARS-CoV-2 pandemic has profoundly impacted individuals of all ages and backgrounds, neonates and school-aged children being vulnerable cohorts resulting from the evolving immunological
[...] Read more.
Reflecting on this Special Issue dedicated to pediatric respiratory viruses, it is evident that the shadow cast by the global SARS-CoV-2 pandemic has profoundly impacted individuals of all ages and backgrounds, neonates and school-aged children being vulnerable cohorts resulting from the evolving immunological profiles and limited exposures to immunity-building experienced during this unprecedented era [...]
Full article
(This article belongs to the Special Issue Pediatric Respiratory Viral Infection)
Open AccessArticle
Regions of Bovine Adenovirus-3 Protein VII Involved in Interactions with Viral and Cellular Proteins
by
Shermila Kulanayake, Faryal Dar and Suresh K. Tikoo
Viruses 2024, 16(5), 732; https://doi.org/10.3390/v16050732 - 5 May 2024
Abstract
The L 1 region of bovine adenovirus (BAdV)-3 encodes a multifunctional protein named protein VII. Anti-protein VII sera detected a protein of 26 kDa in transfected or BAdV-3-infected cells, which localizes to nucleus and nucleolus of infected/transfected cells. Analysis of mutant protein VII
[...] Read more.
The L 1 region of bovine adenovirus (BAdV)-3 encodes a multifunctional protein named protein VII. Anti-protein VII sera detected a protein of 26 kDa in transfected or BAdV-3-infected cells, which localizes to nucleus and nucleolus of infected/transfected cells. Analysis of mutant protein VII identified four redundant overlapping nuclear/nucleolar localization signals as deletion of all four potential nuclear/nucleolar localization signals localizes protein VII predominantly to the cytoplasm. The nuclear import of protein VII appears to use importin α (α-1), importin-β (β-1) and transportin-3 nuclear transport receptors. In addition, different nuclear transport receptors also require part of protein VII outside nuclear localization sequences for efficient interaction. Proteomic analysis of protein complexes purified from recombinant BAdV-3 expressing protein VII containing Strep Tag II identified potential viral and cellular proteins interacting with protein VII. Here, we confirm that protein VII interacts with IVa2 and protein VIII in BAdV-3-infected cells. Moreover, amino acids 91–101 and 126–137, parts of non-conserved region of protein VII, are required for interaction with IVa2 and protein VIII, respectively.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures

Figure 1
Open AccessBrief Report
Rapid Detection and Quick Characterization of African Swine Fever Virus Using the VolTRAX Automated Library Preparation Platform
by
Vivian O’Donnell, Jim L. Pierce, Oleg Osipenko, Lizhe Xu, Amy Berninger, Steven M. Lakin, Roger W. Barrette, Douglas P. Gladue and Bonto Faburay
Viruses 2024, 16(5), 731; https://doi.org/10.3390/v16050731 - 5 May 2024
Abstract
African swine fever virus (ASFV) is the causative agent of a severe and highly contagious viral disease affecting domestic and wild swine. The current ASFV pandemic strain has a high mortality rate, severely impacting pig production and, for countries suffering outbreaks, preventing the
[...] Read more.
African swine fever virus (ASFV) is the causative agent of a severe and highly contagious viral disease affecting domestic and wild swine. The current ASFV pandemic strain has a high mortality rate, severely impacting pig production and, for countries suffering outbreaks, preventing the export of their pig products for international trade. Early detection and diagnosis of ASFV is necessary to control new outbreaks before the disease spreads rapidly. One of the rate-limiting steps to identify ASFV by next-generation sequencing platforms is library preparation. Here, we investigated the capability of the Oxford Nanopore Technologies’ VolTRAX platform for automated DNA library preparation with downstream sequencing on Nanopore sequencing platforms as a proof-of-concept study to rapidly identify the strain of ASFV. Within minutes, DNA libraries prepared using VolTRAX generated near-full genome sequences of ASFV. Thus, our data highlight the use of the VolTRAX as a platform for automated library preparation, coupled with sequencing on the MinION Mk1C for field sequencing or GridION within a laboratory setting. These results suggest a proof-of-concept study that VolTRAX is an effective tool for library preparation that can be used for the rapid and real-time detection of ASFV.
Full article
(This article belongs to the Special Issue Early Diagnosis and Surveillance of Transboundary and Emerging Viral Diseases of Animals)
►▼
Show Figures

Figure 1
Open AccessArticle
Regional Variation of the CD4 and CD8 T Cell Epitopes Conserved in Circulating Dengue Viruses and Shared with Potential Vaccine Candidates
by
Yadya M. Chawla, Prashant Bajpai, Keshav Saini, Elluri Seetharami Reddy, Ashok Kumar Patel, Kaja Murali-Krishna and Anmol Chandele
Viruses 2024, 16(5), 730; https://doi.org/10.3390/v16050730 - 5 May 2024
Abstract
As dengue expands globally and many vaccines are under trials, there is a growing recognition of the need for assessing T cell immunity in addition to assessing the functions of neutralizing antibodies during these endeavors. While several dengue-specific experimentally validated T cell epitopes
[...] Read more.
As dengue expands globally and many vaccines are under trials, there is a growing recognition of the need for assessing T cell immunity in addition to assessing the functions of neutralizing antibodies during these endeavors. While several dengue-specific experimentally validated T cell epitopes are known, less is understood about which of these epitopes are conserved among circulating dengue viruses and also shared by potential vaccine candidates. As India emerges as the epicenter of the dengue disease burden and vaccine trials commence in this region, we have here aligned known dengue specific T cell epitopes, reported from other parts of the world with published polyprotein sequences of 107 dengue virus isolates available from India. Of the 1305 CD4 and 584 CD8 epitopes, we found that 24% and 41%, respectively, were conserved universally, whereas 27% and 13% were absent in any viral isolates. With these data, we catalogued epitopes conserved in circulating dengue viruses from India and matched them with each of the six vaccine candidates under consideration (TV003, TDEN, DPIV, CYD-TDV, DENVax and TVDV). Similar analyses with viruses from Thailand, Brazil and Mexico revealed regional overlaps and variations in these patterns. Thus, our study provides detailed and nuanced insights into regional variation that should be considered for itemization of T cell responses during dengue natural infection and vaccine design, testing and evaluation.
Full article
(This article belongs to the Section Viral Immunology, Vaccines, and Antivirals)
►▼
Show Figures

Graphical abstract

Journal Menu
► ▼ Journal Menu-
- Viruses Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Diseases, Infectious Disease Reports, Pathogens, Viruses, TropicalMed
Human Monkeypox Research
Topic Editors: Shailendra K. Saxena, Ahmed Sayed Abdel-MoneimDeadline: 30 June 2024
Topic in
Biomedicines, JCM, Pathogens, Vaccines, Viruses
Discovery and Development of Monkeypox Disease Treatments
Topic Editors: Mohd Imran, Ali A. RabaanDeadline: 31 August 2024
Topic in
Brain Sciences, Clinics and Practice, COVID, Life, Vaccines, Viruses
Multifaceted Efforts from Basic Research to Clinical Practice in Controlling COVID-19 Disease
Topic Editors: Yih-Horng Shiao, Rashi OjhaDeadline: 30 September 2024

Conferences
Special Issues
Special Issue in
Viruses
Animal Coronaviruses: Infection, Prevention, and Antivirals
Guest Editor: Tomomi TakanoDeadline: 20 May 2024
Special Issue in
Viruses
Bacteriophages and Biofilms 2.0
Guest Editors: Zuzanna Drulis-Kawa, Tomasz OlszakDeadline: 31 May 2024
Special Issue in
Viruses
The Inflammasomes - Key Players in Antiviral Response
Guest Editors: Dong-Yan Jin, Tsan Sam XiaoDeadline: 15 June 2024
Special Issue in
Viruses
State-of-the-Art Virology Research in Australia
Guest Editors: Timothy Newsome, Barry SlobedmanDeadline: 30 June 2024
Topical Collections
Topical Collection in
Viruses
Poxviruses
Collection Editors: Giliane de Souza Trindade, Galileu Barbosa Costa, Flavio Guimaraes da Fonseca
Topical Collection in
Viruses
Phage Therapy
Collection Editors: Nina Chanishvili, Jean-Paul Pirnay, Mikael Skurnik
Topical Collection in
Viruses
Coronaviruses
Collection Editors: Luis Martinez-Sobrido, Fernando Almazan Toral
Topical Collection in
Viruses
SARS-CoV-2 and COVID-19
Collection Editors: Luis Martinez-Sobrido, Fernando Almazan Toral








