-
 Rapid Isolation of Phages for the Treatment of Antibiotic Resistant Infections
Rapid Isolation of Phages for the Treatment of Antibiotic Resistant Infections -
 Hospital-Acquired Pneumonia and Ventilator-Associated Pneumonia: A Literature Review
Hospital-Acquired Pneumonia and Ventilator-Associated Pneumonia: A Literature Review -
 Streptococcus pyogenes, the Master of Enzymatic Immunomodulation
Streptococcus pyogenes, the Master of Enzymatic Immunomodulation -
 Animal and In Vitro Models as Powerful Tools to Decipher the Effects of Enteric Pathogens on the Human Gut Microbiota
Animal and In Vitro Models as Powerful Tools to Decipher the Effects of Enteric Pathogens on the Human Gut Microbiota -
 10-Year Experience of Pediatric Autoimmune Neuropsychiatric Disorders Associated with Streptococcal Infections (PANDAS)
10-Year Experience of Pediatric Autoimmune Neuropsychiatric Disorders Associated with Streptococcal Infections (PANDAS)
Journal Description
Microorganisms
Microorganisms
is a scientific, peer-reviewed, open access journal of microbiology, published monthly online by MDPI. The Hellenic Society Mikrobiokosmos (MBK), the Spanish Society for Nitrogen Fixation (SEFIN) and the Society for Microbial Ecology and Disease (SOMED) are affiliated with the Microorganisms, and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q2 (Microbiology (medical))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.1 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about the Microorganisms.
- Companion journal: Applied Microbiology.
Impact Factor:
4.5 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
Growth and Cell Size of Microalga Auxenochlorella protothecoides AS-1 under Different Trophic Modes
Microorganisms 2024, 12(4), 835; https://doi.org/10.3390/microorganisms12040835 (registering DOI) - 20 Apr 2024
Abstract
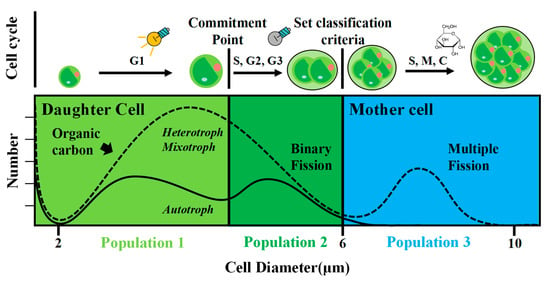
Certain microalgal species can grow with different trophic strategies depending on the availability of nutrient resources. They can use the energy from light or an organic substrate, or both, and can therefore be called autotrophs, heterotrophs, or mixotrophs. We recently isolated a microalgal
[...] Read more.
Certain microalgal species can grow with different trophic strategies depending on the availability of nutrient resources. They can use the energy from light or an organic substrate, or both, and can therefore be called autotrophs, heterotrophs, or mixotrophs. We recently isolated a microalgal strain from the microplastic biofilm, which was identified as Auxenochlorella protothecoides, AS-1. Strain AS-1 grew rapidly in bacterial culture media and exhibited different growth rates and cell sizes under different trophic conditions. We compared the growth performance of AS-1 under the three different trophic modes. AS-1 reached a high biomass (>4 g/L) in 6 days under mixotrophic growth conditions with a few organic carbons as a substrate. In contrast, poor autotrophic growth was observed for AS-1. Different cell sizes, including daughter and mother cells, were observed under the different growth modes. We applied a Coulter Counter to measure the size distribution patterns of AS-1 under different trophic modes. We showed that the cell size distribution of AS-1 was affected by different growth modes. Compared to the auto-, hetero- and mixotrophic modes, AS-1 achieved higher biomass productivity by increasing cell number and cell size in the presence of organic substrate. The mechanisms and advantages of having more mother cells with organic substrates are still unclear and warrant further investigations. The work here provides the growth information of a newly isolated A. protothecoides AS-1 which will be beneficial to future downstream applications.
Full article
(This article belongs to the Section Molecular Microbiology and Immunology)
►
Show Figures
Open AccessArticle
Unveiling Antibacterial Potential and Physiological Characteristics of Thermophilic Bacteria Isolated from a Hot Spring in Iran
by
Zeinab Rafiee, Maryam Jalili Tabaii, Maryam Moradi and Sharareh Harirchi
Microorganisms 2024, 12(4), 834; https://doi.org/10.3390/microorganisms12040834 (registering DOI) - 20 Apr 2024
Abstract
The increasing worldwide demand for antimicrobial agents has significantly contributed to the alarming rise of antimicrobial resistance, posing a grave threat to human life. Consequently, there is a pressing need to explore uncharted environments, seeking out novel antimicrobial compounds that display exceptionally efficient
[...] Read more.
The increasing worldwide demand for antimicrobial agents has significantly contributed to the alarming rise of antimicrobial resistance, posing a grave threat to human life. Consequently, there is a pressing need to explore uncharted environments, seeking out novel antimicrobial compounds that display exceptionally efficient capabilities. Hot springs harbor microorganisms possessing remarkable properties, rendering them an invaluable resource for uncovering groundbreaking antimicrobial compounds. In this study, thermophilic bacteria were isolated from Mahallat Hot Spring, Iran. Out of the 30 isolates examined, 3 strains exhibited the most significant antibacterial activities against Escherichia coli and Staphylococcus aureus. Furthermore, the supernatants of the isolated strains exhibited remarkable antibacterial activity, displaying notable resistance to temperatures as high as 75 °C for 30 min. It was determined that the two strains showed high similarity to the Bacillus genus, while strain Kh3 was classified as Saccharomonospora azurea. All three strains exhibited tolerance to NaCl. Bacillus strains demonstrated optimal growth at pH 5 and 40 °C, whereas S. azurea exhibited optimal growth at pH 9 and 45 °C. Accordingly, hot springs present promising natural reservoirs for the isolation of resilient strains possessing antibacterial properties, which can be utilized in disease treatment or within the food industry.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures

Figure 1
Open AccessReview
Perspectives of FTIR as Promising Tool for Pathogen Diagnosis, Sanitary and Welfare Monitoring in Animal Experimentation Models: A Review Based on Pertinent Literature
by
Matheus Morais Neves, Renan Faria Guerra, Isabela Lima Lemos, Thomas Santos Arrais, Marco Guevara-Vega, Flávia Batista Ferreira, Rafael Borges Rosa, Mylla Spirandelli Vieira, Belchiolina Beatriz Fonseca, Robinson Sabino da Silva and Murilo Vieira da Silva
Microorganisms 2024, 12(4), 833; https://doi.org/10.3390/microorganisms12040833 (registering DOI) - 20 Apr 2024
Abstract
Currently, there is a wide application in the literature of the use of the Fourier Transform Infrared Spectroscopy (FTIR) technique. This basic tool has also proven to be efficient for detecting molecules associated with hosts and pathogens in infections, as well as other
[...] Read more.
Currently, there is a wide application in the literature of the use of the Fourier Transform Infrared Spectroscopy (FTIR) technique. This basic tool has also proven to be efficient for detecting molecules associated with hosts and pathogens in infections, as well as other molecules present in humans and animals’ biological samples. However, there is a crisis in science data reproducibility. This crisis can also be observed in data from experimental animal models (EAMs). When it comes to rodents, a major challenge is to carry out sanitary monitoring, which is currently expensive and requires a large volume of biological samples, generating ethical, legal, and psychological conflicts for professionals and researchers. We carried out a survey of data from the relevant literature on the use of this technique in different diagnostic protocols and combined the data with the aim of presenting the technique as a promising tool for use in EAM. Since FTIR can detect molecules associated with different diseases and has advantages such as the low volume of samples required, low cost, sustainability, and provides diagnostic tests with high specificity and sensitivity, we believe that the technique is highly promising for the sanitary and stress and the detection of molecules of interest of infectious or non-infectious origin.
Full article
(This article belongs to the Section Veterinary Microbiology)
►▼
Show Figures

Figure 1
Open AccessArticle
Priestia megaterium ASC-1 Isolated from Pickled Cabbage Ameliorates Hyperuricemia by Degrading Uric Acid in Rats
by
Wenjuan Zhu, Siyuan Bi, Zhijia Fang, Lukman Iddrisu, Qi Deng, Lijun Sun and Ravi Gooneratne
Microorganisms 2024, 12(4), 832; https://doi.org/10.3390/microorganisms12040832 (registering DOI) - 20 Apr 2024
Abstract
Pickled cabbage, a traditional fermented food rich in functional microorganisms, can effectively control hyperuricemia and gout. In this study, a Priestia megaterium ASC-1 strain with strong uric acid (UA) degradation ability was isolated from pickled cabbage. After oral administration for 15 days, ASC-1
[...] Read more.
Pickled cabbage, a traditional fermented food rich in functional microorganisms, can effectively control hyperuricemia and gout. In this study, a Priestia megaterium ASC-1 strain with strong uric acid (UA) degradation ability was isolated from pickled cabbage. After oral administration for 15 days, ASC-1 was stably colonized in the rats in this study. ASC-1 significantly reduced UA levels (67.24%) in hyperuricemic rats. Additionally, ASC-1 alleviated hyperuricemia-related inflammatory response, oxidative stress, and blood urea nitrogen. Intestinal microbial diversity results showed that ASC-1 restored intestinal injury and gut flora dysbiosis caused by hyperuricemia. These findings suggest that P. megaterium ASC-1 may be used as a therapeutic adjuvant for the treatment of hyperuricemia.
Full article
(This article belongs to the Special Issue Nutritional Regulation on Gut Microbiota, 2nd Edition)
►▼
Show Figures
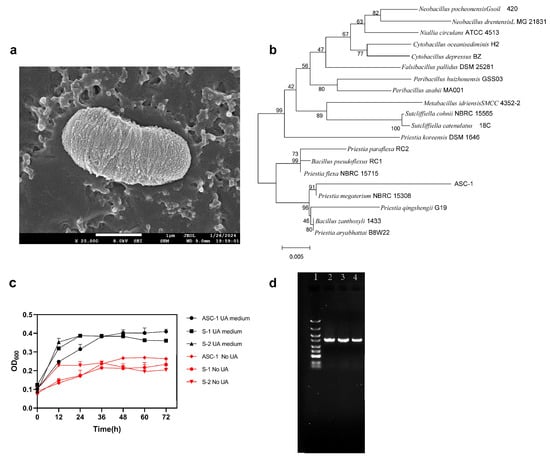
Figure 1
Open AccessArticle
Evaluation of Probiotic Properties and Safety of Lactobacillus helveticus LH10 Derived from Vinegar through Comprehensive Analysis of Genotype and Phenotype
by
Yang Du, Jingru Xu, Jinquan Li and Renwei Wu
Microorganisms 2024, 12(4), 831; https://doi.org/10.3390/microorganisms12040831 (registering DOI) - 19 Apr 2024
Abstract
The probiotic potential of Lactobacillus helveticus LH10, derived from vinegar Pei, a brewing mixture, was assessed through genotype and phenotype analyses. The assembled genome was comprised of 1,810,276 bp and predicted a total of 2044 coding sequences (CDSs). Based on the whole genome
[...] Read more.
The probiotic potential of Lactobacillus helveticus LH10, derived from vinegar Pei, a brewing mixture, was assessed through genotype and phenotype analyses. The assembled genome was comprised of 1,810,276 bp and predicted a total of 2044 coding sequences (CDSs). Based on the whole genome sequence analysis, two bacteriocin gene clusters were identified, while no pathogenic genes were detected. In in vitro experiments, L. helveticus LH10 exhibited excellent tolerance to simulated gastrointestinal fluid, a positive hydrophobic interaction with xylene, and good auto-aggregation properties. Additionally, this strain demonstrated varying degrees of resistance to five antibiotics, strong antagonistic activity against four tested pathogens, and no hemolytic activity. Therefore, L. helveticus LH10 holds great promise as a potential probiotic candidate deserving further investigation for its beneficial effects on human health.
Full article
(This article belongs to the Special Issue Food Microorganisms and Genomics)
Open AccessArticle
Impact of Multidrug-Resistant Organisms on Severe Acquired Brain Injury Rehabilitation: An Observational Study
by
Giovanna Barbara Castellani, Elisa Maietti, Valentina Colombo, Stefano Clemente, Ivo Cassani and Paola Rucci
Microorganisms 2024, 12(4), 830; https://doi.org/10.3390/microorganisms12040830 - 19 Apr 2024
Abstract
Healthcare-associated infections (HAIa) and antimicrobial resistance are expected to be the next threat to human health and are most frequent in people with severe acquired brain injury (SABI), who can be more easily colonized by multidrug-resistant organisms (MDROs). The study’s aim is to
[...] Read more.
Healthcare-associated infections (HAIa) and antimicrobial resistance are expected to be the next threat to human health and are most frequent in people with severe acquired brain injury (SABI), who can be more easily colonized by multidrug-resistant organisms (MDROs). The study’s aim is to investigate the impact of MDRO colonizations and infections on SABI rehabilitation outcomes. This retrospective observational study was performed in a tertiary referral specialized rehabilitation hospital. The main outcomes were the presence of carbapenemase-producing Enterobacteriaceae (CPE) colonization, type and timing of HAI and MDRO HAI, and the number of CPE transmissions. We included 48 patients, 31% carrying CPE on admission and 33% colonized during the hospitalization. A total of 101 HAI were identified in 40 patients, with an overall incidence of 10.5/1000 patient days. Some 37% of patients had at least one MDRO infection, with a MDRO infection incidence of 2.8/1000 patient days. The number of HAIs was significantly correlated with the length of stay (LOS) (r = 0.453, p = 0.001). A significant correlation was found between colonization and type of hospital room (p = 0.013). Complications and HAI significantly affected LOS. We suggest that CPE carriers might be at risk of HAI and worse outcomes compared with non-CPE carriers.
Full article
(This article belongs to the Special Issue Antimicrobial Resistance: Challenges and Innovative Solutions)
Open AccessArticle
Association of Acidotolerant Cyanobacteria to Microbial Mats below pH 1 in Acidic Mineral Precipitates in Río Tinto River in Spain
by
Felipe Gómez, Nuria Rodríguez, José Antonio Rodríguez-Manfredi, Cristina Escudero, Ignacio Carrasco-Ropero, José M. Martínez, Marco Ferrari, Simone De Angelis, Alessandro Frigeri, Maite Fernández-Sampedro and Ricardo Amils
Microorganisms 2024, 12(4), 829; https://doi.org/10.3390/microorganisms12040829 - 19 Apr 2024
Abstract
This report describes acidic microbial mats containing cyanobacteria that are strongly associated to precipitated minerals in the source area of Río Tinto. Río Tinto (Huelva, Southwestern Spain) is an extreme acidic environment where iron and sulfur cycles play a fundamental role in sustaining
[...] Read more.
This report describes acidic microbial mats containing cyanobacteria that are strongly associated to precipitated minerals in the source area of Río Tinto. Río Tinto (Huelva, Southwestern Spain) is an extreme acidic environment where iron and sulfur cycles play a fundamental role in sustaining the extremely low pH and the high concentration of heavy metals, while maintaining a high level of microbial diversity. These multi-layered mineral deposits are stable all year round and are characterized by a succession of thick greenish-blue and brownish layers mainly composed of natrojarosite. The temperature and absorbance above and below the mineral precipitates were followed and stable conditions were detected inside the mineral precipitates. Different methodologies, scanning and transmission electron microscopy, immunological detection, fluorescence in situ hybridization, and metagenomic analysis were used to describe the biodiversity existing in these microbial mats, demonstrating, for the first time, the existence of acid-tolerant cyanobacteria in a hyperacidic environment of below pH 1. Up to 0.46% of the classified sequences belong to cyanobacterial microorganisms, and 1.47% of the aligned DNA reads belong to the Cyanobacteria clade.
Full article
(This article belongs to the Collection Microbial Life in Extreme Environments)
Open AccessArticle
Effect of Stool Sampling on a Routine Clinical Method for the Quantification of Six Short Chain Fatty Acids in Stool Using Gas Chromatography–Mass Spectrometry
by
Tarek Mahdi, Aurore Desmons, Pranvera Krasniqi, Jean-Marc Lacorte, Nathalie Kapel, Antonin Lamazière, Salma Fourati and Thibaut Eguether
Microorganisms 2024, 12(4), 828; https://doi.org/10.3390/microorganisms12040828 - 19 Apr 2024
Abstract
Short chain fatty acids (SCFAs) are primarily produced in the caecum and proximal colon via the bacterial fermentation of undigested carbohydrates that have avoided digestion in the small intestine. Increasing evidence supports the critical role that SCFAs play in health and homeostasis. Microbial
[...] Read more.
Short chain fatty acids (SCFAs) are primarily produced in the caecum and proximal colon via the bacterial fermentation of undigested carbohydrates that have avoided digestion in the small intestine. Increasing evidence supports the critical role that SCFAs play in health and homeostasis. Microbial SCFAs, namely butyric acid, serve as a principal energy source for colonocytes, and their production is essential for gut integrity. A direct link between SCFAs and some human pathological conditions, such as inflammatory bowel disease, irritable bowel syndrome, diarrhea, and cancer, has been proposed. The direct measurement of SCFAs in feces provides a non-invasive approach to demonstrating connections between SCFAs, microbiota, and metabolic diseases to estimate their potential applicability as meaningful biomarkers of intestinal health. This study aimed to adapt a robust analytical method (liquid–liquid extraction, followed by isobutyl chloroformate derivatization and GC–MS analysis), with comparable performances to methods from the literature, and to use this tool to tackle the question of pre-analytical conditions, namely stool processing. We focused on the methodology of managing stool samples before the analysis (fresh stool or dilution in either ethanol/methanol, lyophilized stool, or RNAlater®), as this is a significant issue to consider for standardizing results between clinical laboratories. The objective was to standardize methods for future applications as diagnostic tools. In this paper, we propose a validated GC–MS method for SCFA quantification in stool samples, including pre- and post-analytical comparison studies that could be easily used for clinical laboratory purposes. Our results show that using lyophilization as a stool-processing method would be the best method to achieve this goal.
Full article
(This article belongs to the Section Gut Microbiota)
Open AccessReview
Neutrophils versus Protozoan Parasites: Plasmodium, Trichomonas, Leishmania, Trypanosoma, and Entameoba
by
Eileen Uribe-Querol and Carlos Rosales
Microorganisms 2024, 12(4), 827; https://doi.org/10.3390/microorganisms12040827 - 19 Apr 2024
Abstract
Neutrophils are the most abundant polymorphonuclear granular leukocytes in human blood and are an essential part of the innate immune system. Neutrophils are efficient cells that eliminate pathogenic bacteria and fungi, but their role in dealing with protozoan parasitic infections remains controversial. At
[...] Read more.
Neutrophils are the most abundant polymorphonuclear granular leukocytes in human blood and are an essential part of the innate immune system. Neutrophils are efficient cells that eliminate pathogenic bacteria and fungi, but their role in dealing with protozoan parasitic infections remains controversial. At sites of protozoan parasite infections, a large number of infiltrating neutrophils is observed, suggesting that neutrophils are important cells for controlling the infection. Yet, in most cases, there is also a strong inflammatory response that can provoke tissue damage. Diseases like malaria, trichomoniasis, leishmaniasis, Chagas disease, and amoebiasis affect millions of people globally. In this review, we summarize these protozoan diseases and describe the novel view on how neutrophils are involved in protection from these parasites. Also, we present recent evidence that neutrophils play a double role in these infections participating both in control of the parasite and in the pathogenesis of the disease.
Full article
(This article belongs to the Special Issue Current Insights into Host–Parasite Interactions)
►▼
Show Figures
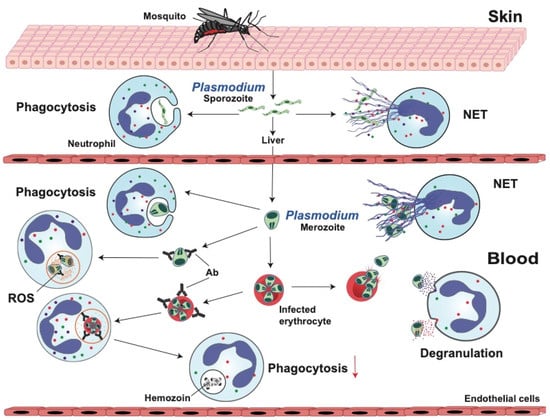
Figure 1
Open AccessArticle
Acute Hepatitis of Unknown Origin in Children: Analysis of 17 Cases Admitted to the Bambino Gesù Children’s Hospital in Rome
by
Velia Chiara Di Maio, Leonarda Gentile, Rossana Scutari, Luna Colagrossi, Luana Coltella, Stefania Ranno, Giulia Linardos, Daniela Liccardo, Maria Sole Basso, Andrea Pietrobattista, Simona Landi, Lorena Forqué, Marta Ciofi Degli Atti, Lara Ricotta, Andrea Onetti Muda, Giuseppe Maggiore, Massimiliano Raponi, Carlo Federico Perno and Cristina Russo
Microorganisms 2024, 12(4), 826; https://doi.org/10.3390/microorganisms12040826 - 19 Apr 2024
Abstract
This study described 17 cases of children admitted to the Bambino Gesù Children’s Hospital with acute hepatitis of unknown origin between mid-April and November 2022. Following the World Health Organization’s working case definition of probable cases, 17 children, with a median age of
[...] Read more.
This study described 17 cases of children admitted to the Bambino Gesù Children’s Hospital with acute hepatitis of unknown origin between mid-April and November 2022. Following the World Health Organization’s working case definition of probable cases, 17 children, with a median age of 2.1 years (interquartile range: 1.0–7.1), presenting with acute hepatitis non-AE, with serum transaminase >500 IU/L, were included in the study. A pre-specified set of microbiological tests was performed on different biological specimens for all pediatric patients. All patients resulted negative for the common hepatotropic viruses. The most common pathogen detected in blood specimens was human-herpes-virus-7 (52.9%). Adenovirus was detected more frequently in stool specimens (62.5%) than in respiratory (20.0%) or blood samples (17.6%). Regarding Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) infection, one child tested positive two days after admission, while antibodies against spike and nucleoprotein were present in 82.3% of patients. A co-pathogen detection was observed in 94.1% of children. Overall, 16 children recovered without clinical complications, while one patient required liver transplantation. In these cases of acute hepatitis of unknown origin, adenovirus was mainly detected in stool samples. A co-pathogen detection was also frequently observed, suggesting that the etiology of this acute hepatitis is most probably multifactorial.
Full article
(This article belongs to the Special Issue Recent Advances in Antivirals for Emerging Viruses 3.0)
►▼
Show Figures
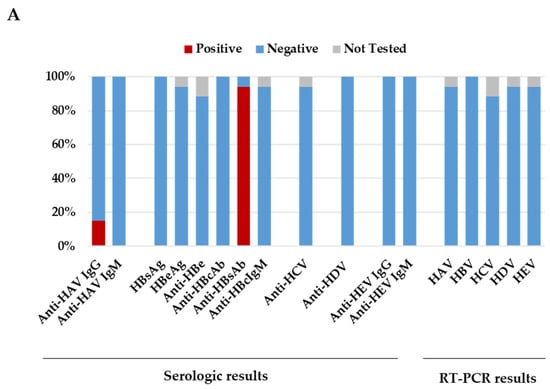
Figure 1
Open AccessArticle
Evaluation of the Effects of Food Safety Training on the Microbiological Load Present in Equipment, Surfaces, Utensils, and Food Manipulator’s Hands in Restaurants
by
Miguel Castro, Kamila Soares, Carlos Ribeiro and Alexandra Esteves
Microorganisms 2024, 12(4), 825; https://doi.org/10.3390/microorganisms12040825 - 19 Apr 2024
Abstract
Training food handlers is essential to ensure food safety. However, the efficacy of training programs relying solely on theoretical information remains uncertain and often fails to induce significant changes in inadequate food practices. Training programs in good hygiene and food safety practices that
[...] Read more.
Training food handlers is essential to ensure food safety. However, the efficacy of training programs relying solely on theoretical information remains uncertain and often fails to induce significant changes in inadequate food practices. Training programs in good hygiene and food safety practices that integrate theoretical and practical approaches have emerged as a vital tool, enabling food handlers to apply their knowledge during work hours and clarify doubts. This study aimed to assess the impact of food safety training based on theoretical and on-the-job training on the microbiological counts of equipment, surfaces, utensils, and food handler (FH) hands. The hygiene and food safety conditions of four restaurants were analyzed through facility checklists, employee questionnaires, and microbiological analyses conducted before and after training. Eight sample collection moments were conducted at each restaurant before and after training. The pre-training results indicate that 15% and 26% of analyses for Enterobacteriaceae and total mesophilic aerobic bacteria (TMB), respectively, did not comply with hygiene safety limits. Additionally, 31% and 64% of Enterobacteriaceae and TMB values, respectively, exceeded safety limits on food handler hands. Positive cases of coagulase-positive Staphylococcus (CoPS) resulted from unprotected wounds on some FH hands. The presence of Listeria monocytogenes in drains was also identified as a concern. Following training, significant differences in results were observed. In many cases, there was a reduction of over 80% in microbial load for Enterobacteriaceae and TMB collected from equipment, surfaces, utensils, and food handler hands. The presence of L. monocytogenes in drains was also eliminated after food safety training. In conclusion, this study underscores the importance of effective training in improving food safety practices.
Full article
(This article belongs to the Special Issue Selected Papers from the 2nd International Electronic Conference on Microbiology (ECM 2023))
►▼
Show Figures
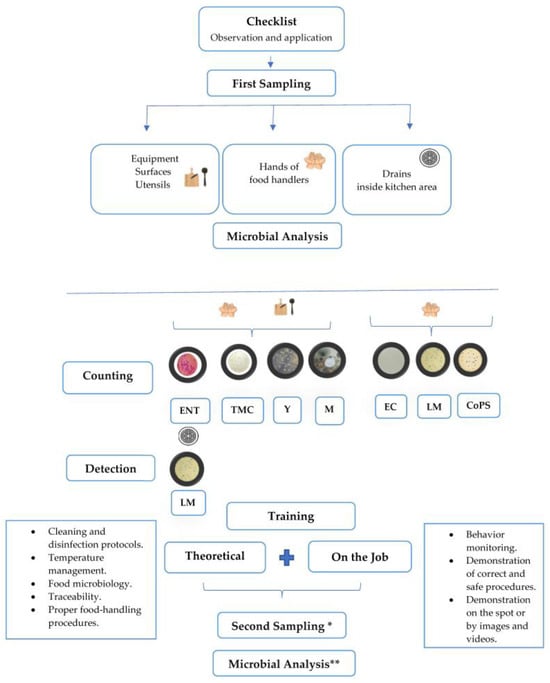
Figure 1
Open AccessArticle
Whole-Genome Sequencing and Phenotypic Analysis of Streptococcus equi subsp. zooepidemicus Sequence Type 147 Isolated from China
by
Yan Su, Zehua Zhang, Li Wang, Baojiang Zhang and Lingling Su
Microorganisms 2024, 12(4), 824; https://doi.org/10.3390/microorganisms12040824 - 19 Apr 2024
Abstract
Streptococcus equi subsp. zooepidemicus (S. zooepidemicus) is one of the important zoonotic and opportunistic pathogens. In recent years, there has been growing evidence that supports the potential role of S. zooepidemicus in severe diseases in horses and other animals, including humans.
[...] Read more.
Streptococcus equi subsp. zooepidemicus (S. zooepidemicus) is one of the important zoonotic and opportunistic pathogens. In recent years, there has been growing evidence that supports the potential role of S. zooepidemicus in severe diseases in horses and other animals, including humans. Furthermore, the clinical isolation and drug resistance rates of S. zooepidemicus have been increasing yearly, leading to interest in its in-depth genomic analysis. In order to deepen the understanding of the S. zooepidemicus characteristics and genomic features, we investigated the genomic islands, mobile genetic elements, virulence and resistance genes, and phenotype of S. zooepidemicus strain ZHZ 211 (ST147), isolated from an equine farm in China. We obtained a 2.18 Mb, high-quality chromosome and found eight genomic islands. According to a comparative genomic investigation with other reference strains, ZHZ 211 has more virulence factors, like an iron uptake system, adherence, exoenzymes, and antiphagocytosis. More interestingly, ZHZ 211 has acquired a mobile genetic element (MGE), prophage Ph01, which was found to be in the chromosome of this strain and included two hyaluronidase (hyl) genes, important virulence factors of the strain. Moreover, two transposons and two virulence (virD4) genes were found to be located in the same genome island of ZHZ 211. In vitro phenotypic results showed that ZHZ 211 grows faster and is resistant to clarithromycin, enrofloxacin, and sulfonamides. The higher biofilm-forming capabilities of ZHZ 211 may provide a competitive advantage for survival in its niche. The results expand our understanding of the genomic, pathogenicity, and resistance characterization of Streptococcus zooepidemicus and facilitate further exploration of its molecular pathogenic mechanism.
Full article
(This article belongs to the Section Veterinary Microbiology)
►▼
Show Figures
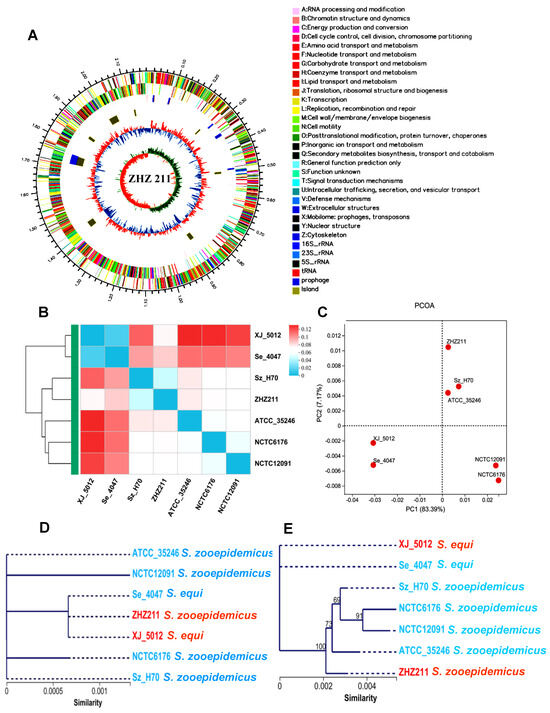
Figure 1
Open AccessArticle
Comparative Analysis of Bisexual and Parthenogenetic Populations in Haemaphysalis Longicornis
by
Chaoyue Zhao, Guonan Cai, Xing Zhang, Xinyu Liu, Pengfei Wang and Aihua Zheng
Microorganisms 2024, 12(4), 823; https://doi.org/10.3390/microorganisms12040823 - 19 Apr 2024
Abstract
Haemaphysalis longicornis, a three-host tick with a wide host range, is widely distributed in different countries and regions. It stands out among ticks due to its unique feature of having both parthenogenetic and bisexual populations. Despite their morphological resemblance, the characteristics of
[...] Read more.
Haemaphysalis longicornis, a three-host tick with a wide host range, is widely distributed in different countries and regions. It stands out among ticks due to its unique feature of having both parthenogenetic and bisexual populations. Despite their morphological resemblance, the characteristics of the parthenogenetic population have been overlooked. In this comprehensive study, we systematically compared the similarities and differences between these two populations. Our investigation revealed that the parthenogenetic H. longicornis, widely distributed in China, was found in ten provinces, surpassing the previously reported distribution. Notably, individuals from the parthenogenetic population exhibited a prolonged blood-feeding duration during the larval and nymph stages compared to their bisexual counterparts. Additionally, the life cycle of the parthenogenetic population was observed to be longer. A flow cytometry analysis indicated a DNA content ratio of approximately 2:3 between the bisexual and parthenogenetic populations. A phylogenetic analysis using whole mitochondrial genome sequences resulted in the separation of the phylogenetic tree into two distinct branches. A molecular analysis unveiled a consistent single T-base deletion at nucleotide 8497 in the parthenogenetic population compared to the bisexual population. Both populations displayed high viral infection capability and significant resistance to ivermectin. Intriguingly, despite these differences, the parthenogenetic population exhibited a similar life cycle to the bisexual population, retaining the ability to transmit pathogens such as Severe fever with thrombocytopenia syndrome virus (SFTSV) and Heartland Virus (HRTV). These findings contribute to a deeper understanding of the distinct characteristics and similarities between different populations of H. longicornis, laying the foundation for future research in this field.
Full article
(This article belongs to the Section Parasitology)
►▼
Show Figures
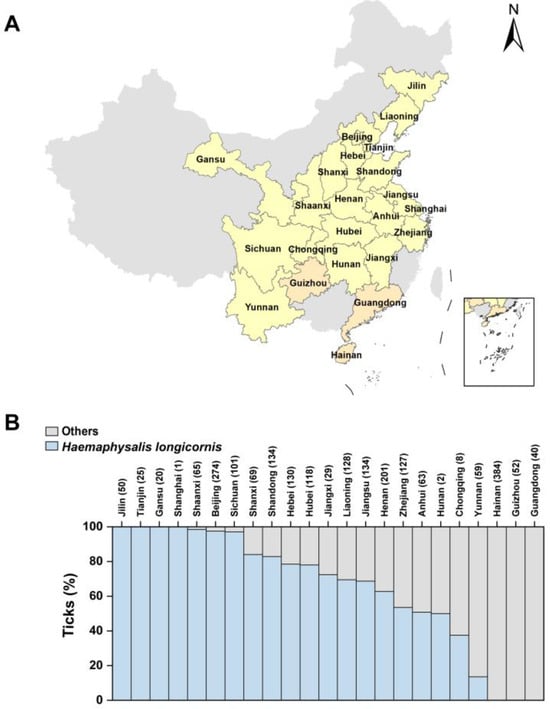
Figure 1
Open AccessArticle
The Changes in Fecal Bacterial Communities in Goats Offered Rumen-Protected Fat
by
Hu Liu, Weishi Peng, Kaiyu Mao, Yuanting Yang, Qun Wu, Ke Wang, Meng Zeng, Xiaotao Han, Jiancheng Han and Hanlin Zhou
Microorganisms 2024, 12(4), 822; https://doi.org/10.3390/microorganisms12040822 - 19 Apr 2024
Abstract
Leizhou goats are famous for their delicious meat but have inferior growth performance. There is little information on rumen-protected fat (RPF) from the Leizhou goat. Hence, we observed the effects of RPF on growth, fecal short-chain fatty acids, and bacteria community with respect
[...] Read more.
Leizhou goats are famous for their delicious meat but have inferior growth performance. There is little information on rumen-protected fat (RPF) from the Leizhou goat. Hence, we observed the effects of RPF on growth, fecal short-chain fatty acids, and bacteria community with respect to Leizhou goats. Twelve goats (13.34 ± 0.024 kg) were selected and assigned randomly to one of two treatments: (1) a control diet (CON) and (2) 2.4% RPF with a control diet (RPF). The final body weight and average daily gain (ADG) were greater (p < 0.05), and the dry matter intake (DMI): ADG was lower (p < 0.05) in the RPF group than in the CON group. There were no differences in DMI between the CON and RPF groups. The concentrations of total short-chain fatty acids, acetate, propionate, and butyrate were lower (p < 0.05) in the RPF group than in the CON group. The relative abundances of Ruminococcus, Rikenellaceae_RC9_gut_group, Treponema, norank_f__norank_o__RF39, Eubacterium_siraeum_group, and Ruminococcus_torques_group were lower (p < 0.05) in the RPF group than in the CON group. The relative abundances of Bacteroides, norank_f__norank_o__Clostridia_UCG-014, norank_f__Eubacterium_coprostanoligenes_group, Eubacterium_ruminantium_group, norank_f__Oscillospirale-UCG-010, Oscillospiraceae_UCG-002, and Family_XIII_AD3011_group were greater (p < 0.05) in the RPF group than in the CON group. It was concluded that RPF could improve the goats’ growth performance by regulating their fecal bacteria communities.
Full article
(This article belongs to the Section Veterinary Microbiology)
►▼
Show Figures
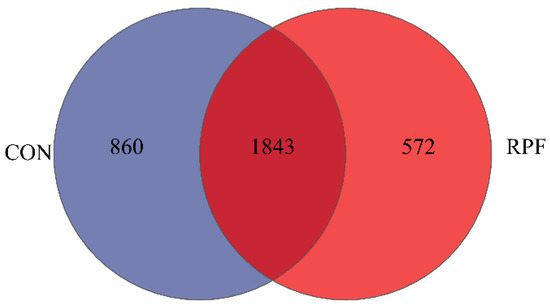
Figure 1
Open AccessArticle
Sex- and Gender-Based Analysis on Norepinephrine Use in Septic Shock: Why Is It Still a Male World?
by
Benedetta Perna, Valeria Raparelli, Federica Tordo Caprioli, Oana Teodora Blanaru, Cecilia Malacarne, Cecilia Crosetti, Andrea Portoraro, Alex Zanotto, Francesco Maria Strocchi, Alessandro Rapino, Anna Costanzini, Martina Maritati, Roberto Lazzari, Michele Domenico Spampinato, Carlo Contini, Roberto De Giorgio and Matteo Guarino
Microorganisms 2024, 12(4), 821; https://doi.org/10.3390/microorganisms12040821 - 18 Apr 2024
Abstract
Sex and gender are fundamental health determinants and their role as modifiers of treatment response is increasingly recognized. Norepinephrine is a cornerstone of septic shock management and its use is based on the highest level of evidence compared to dopamine. The related 2021
[...] Read more.
Sex and gender are fundamental health determinants and their role as modifiers of treatment response is increasingly recognized. Norepinephrine is a cornerstone of septic shock management and its use is based on the highest level of evidence compared to dopamine. The related 2021 Surviving Sepsis Campaign (SCC) recommendation is presumably applicable to both females and males; however, a sex- and gender-based analysis is lacking, thus not allowing generalizable conclusions. This paper was aimed at exploring whether sex- and gender-disaggregated data are available in the evidence supporting this recommendation. For all the studies underpinning it, four pairs of authors, including a woman and a man, extracted data concerning sex and gender, according to the Sex and Gender Equity in Research guidelines. Nine manuscripts were included with an overall population of 2126 patients, of which 43.2% were females. No sex analysis was performed and gender was never reported. In conclusion, the present manuscript highlighted that the clinical studies underlying the SCC recommendation of NE administration in septic shock have neglected the likely role of sex and gender as modifiers of treatment response, thus missing the opportunity of sex- and gender-specific guidelines.
Full article
(This article belongs to the Special Issue Overview of Sepsis and Septic Shock)
►▼
Show Figures

Figure 1
Open AccessArticle
Ligand-Free Silver Nanoparticles: An Innovative Strategy against Viruses and Bacteria
by
Maria Vittoria Morone, Annalisa Chianese, Federica Dell’Annunziata, Veronica Folliero, Erwin Pavel Lamparelli, Giovanna Della Porta, Carla Zannella, Anna De Filippis, Gianluigi Franci, Massimiliano Galdiero and Antonio Morone
Microorganisms 2024, 12(4), 820; https://doi.org/10.3390/microorganisms12040820 - 18 Apr 2024
Abstract
The spread of antibiotic-resistant bacteria and the rise of emerging and re-emerging viruses in recent years constitute significant public health problems. Therefore, it is necessary to develop new antimicrobial strategies to overcome these challenges. Herein, we describe an innovative method to synthesize ligand-free
[...] Read more.
The spread of antibiotic-resistant bacteria and the rise of emerging and re-emerging viruses in recent years constitute significant public health problems. Therefore, it is necessary to develop new antimicrobial strategies to overcome these challenges. Herein, we describe an innovative method to synthesize ligand-free silver nanoparticles by Pulsed Laser Ablation in Liquid (PLAL-AgNPs). Thus produced, nanoparticles were characterized by total X-ray fluorescence, zeta potential analysis, transmission electron microscopy (TEM), and nanoparticle tracking analysis (NTA). A 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay was performed to evaluate the nanoparticles’ cytotoxicity. Their potential was evaluated against the enveloped herpes simplex virus type 1 (HSV-1) and the naked poliovirus type 1 (PV-1) by plaque reduction assays and confirmed by real-time PCR and fluorescence microscopy, showing that nanoparticles interfered with the early stage of infection. Their action was also examined against different bacteria. We observed that the PLAL-AgNPs exerted a strong effect against both methicillin-resistant Staphylococcus aureus (S. aureus MRSA) and Escherichia coli (E. coli) producing extended-spectrum β-lactamase (ESBL). In detail, the PLAL-AgNPs exhibited a bacteriostatic action against S. aureus and a bactericidal activity against E. coli. Finally, we proved that the PLAL-AgNPs were able to inhibit/degrade the biofilm of S. aureus and E. coli.
Full article
(This article belongs to the Special Issue Antimicrobial Properties of Nanoparticle)
►▼
Show Figures
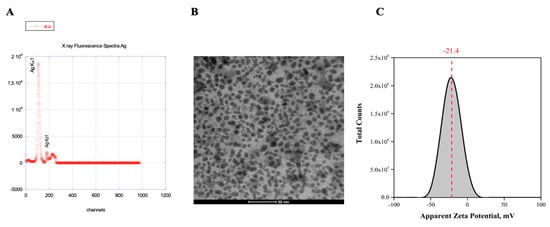

Figure 1
Open AccessArticle
Epidemiological and Clinical Aspects of Cutaneous and Mucosal Leishmaniases in Portugal: Retrospective Analysis of Cases Diagnosed in Public Hospitals and Reported in the Literature between 2010 and 2020
by
Rafael Rocha, Cláudia Conceição, Luzia Gonçalves, Ana Cláudia Carvalho, André Maia, André Martins, António Carujo, António Maio, Catarina Forra, Catarina Melita, Daniela Couto, Diana Fernandes, Dulce Pereira, Ema Leal, Helena Sarmento, Inês Sousa, Jean-Pierre Gonçalves, Joana Marinho, Joana Vasconcelos, João Cunha, João Rodrigues, José Miguel Silva, Lídia Caley, Luís Malheiro, Luís Santos, Margarida Garcia, Maria Cunha, Maria Lima, Maria Margarida Andrade, Marta Marques, Miguel Alpalhão, Mónica Silva, Rita Ferraz, Rui Soares, Salomão Fernandes, Samuel Llobet, Sofia Cruz, Teresa Guimarães, Tiago Branco, Tomás Robalo-Nunes, Vasco Almeida and Carla Maiaadd
Show full author list
remove
Hide full author list
Microorganisms 2024, 12(4), 819; https://doi.org/10.3390/microorganisms12040819 - 18 Apr 2024
Abstract
Leishmania infantum, a zoonotic vector-born parasite, is endemic in the Mediterranean region, presenting mostly as visceral (VL), but also as cutaneous (CL) and mucosal leishmaniasis (ML). This study aimed to describe the epidemiological and clinical aspects of the CL and ML cases
[...] Read more.
Leishmania infantum, a zoonotic vector-born parasite, is endemic in the Mediterranean region, presenting mostly as visceral (VL), but also as cutaneous (CL) and mucosal leishmaniasis (ML). This study aimed to describe the epidemiological and clinical aspects of the CL and ML cases diagnosed in mainland Portugal between 2010 and 2020. Collaboration was requested from every hospital of the Portuguese National Health System. Cases were screened through a search of diagnostic discharge codes or positive laboratory results for Leishmania infection. Simultaneously, a comprehensive literature search was performed. Descriptive statistics and hypothesis testing were performed using IBM® SPSS® Statistics. A total of 43 CL and 7 ML cases were identified, with a predominance of autochthonous cases (86%). In CL, immunosuppressed individuals constituted a significant proportion of patients (48%), and in this group, disseminated CL (22%) and simultaneous VL (54%) were common. In autochthonous cases, lesions, mostly papules/nodules (62%), were frequently observed on the head (48%). The approach to treatment was very heterogeneous. ML cases were all autochthonous, were diagnosed primarily in older immunosuppressed individuals, and were generally treated with liposomal amphotericin B. The findings suggest a need for enhanced surveillance and reporting, clinical awareness, and diagnostic capacity of these forms of leishmaniasis to mitigate underdiagnosis and improve patient outcomes. A holistic One Health approach is advocated to address the multifaceted challenges posed by leishmaniases in Portugal and beyond.
Full article
(This article belongs to the Section Parasitology)
►▼
Show Figures
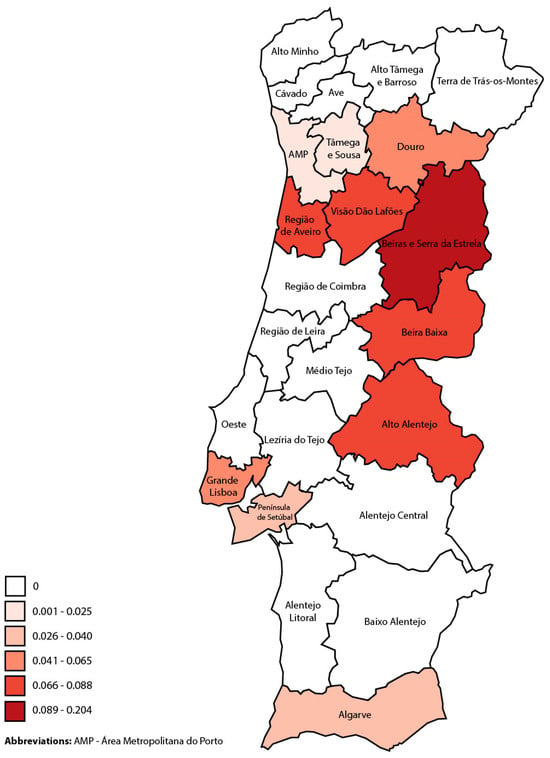
Figure 1
Open AccessReview
Non-O1/Non-O139 Vibrio cholerae—An Underestimated Foodborne Pathogen? An Overview of Its Virulence Genes and Regulatory Systems Involved in Pathogenesis
by
Quantao Zhang, Thomas Alter and Susanne Fleischmann
Microorganisms 2024, 12(4), 818; https://doi.org/10.3390/microorganisms12040818 - 18 Apr 2024
Abstract
In recent years, the number of foodborne infections with non-O1 and non-O139 Vibrio cholerae (NOVC) has increased worldwide. These have ranged from sporadic infection cases to localized outbreaks. The majority of case reports describe self-limiting gastroenteritis. However, severe gastroenteritis and even cholera-like symptoms
[...] Read more.
In recent years, the number of foodborne infections with non-O1 and non-O139 Vibrio cholerae (NOVC) has increased worldwide. These have ranged from sporadic infection cases to localized outbreaks. The majority of case reports describe self-limiting gastroenteritis. However, severe gastroenteritis and even cholera-like symptoms have also been described. All reported diarrheal cases can be traced back to the consumption of contaminated seafood. As climate change alters the habitats and distribution patterns of aquatic bacteria, there is a possibility that the number of infections and outbreaks caused by Vibrio spp. will further increase, especially in countries where raw or undercooked seafood is consumed or clean drinking water is lacking. Against this background, this review article focuses on a possible infection pathway and how NOVC can survive in the human host after oral ingestion, colonize intestinal epithelial cells, express virulence factors causing diarrhea, and is excreted by the human host to return to the environment.
Full article
(This article belongs to the Special Issue Vibrio Virulence)
►▼
Show Figures
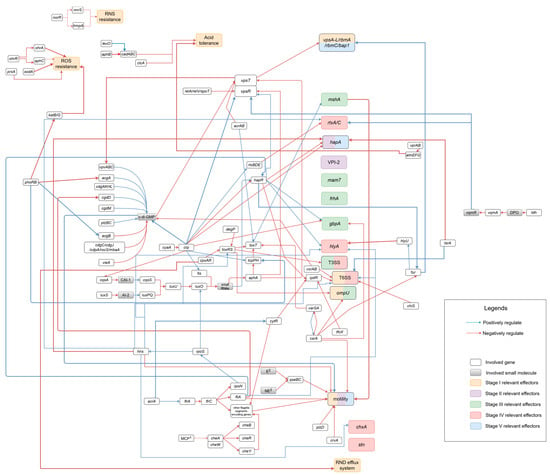
Figure 1
Open AccessArticle
Genomic Characterization of Listeria monocytogenes and Other Listeria Species Isolated from Sea Turtles
by
Ludovica Di Renzo, Maria Elisabetta De Angelis, Marina Torresi, Giulia Mariani, Federica Pizzurro, Luana Fiorella Mincarelli, Emanuele Esposito, Maria Oliviero, Doriana Iaccarino, Fabio Di Nocera, Gianluigi Paduano, Giuseppe Lucifora, Cesare Cammà, Nicola Ferri and Francesco Pomilio
Microorganisms 2024, 12(4), 817; https://doi.org/10.3390/microorganisms12040817 - 18 Apr 2024
Abstract
Listeria monocytogenes is a ubiquitous pathogen found both in the environment and food. It can cause listeriosis in a wide range of animals as well as in humans. Investigations on presence, spread and virulence are still limited to terrestrial and human environments. Embracing
[...] Read more.
Listeria monocytogenes is a ubiquitous pathogen found both in the environment and food. It can cause listeriosis in a wide range of animals as well as in humans. Investigations on presence, spread and virulence are still limited to terrestrial and human environments. Embracing the One Health Approach, investigating the presence and spread of L. monocytogenes in marine ecosystems and among wildlife, would provide us with useful information for human health. This study investigated the presence of L. monocytogenes and Listeria spp. in two species of sea turtles common in the Mediterranean Sea (Caretta caretta and Chelonia mydas). A total of one hundred and sixty-four carcasses of sea turtles (C. caretta n = 161 and C. mydas n = 3) stranded along the Abruzzo, Molise, Campania, and Calabria coasts, were collected. Brain and fecal samples were taken, enriched, and cultured for the detection of Listeria spp. From the specimens collected, strains of L. monocytogenes (brain n = 1, brain and feces n = 1, multiorgan n = 1 and feces n = 1), L. innocua (feces n = 1 and brain n = 1), and L. ivanovii (brain n = 1) were isolated. Typical colonies were isolated for Whole Genome Sequencing (WGS). Virulence genes, disinfectants/metal resistance, and antimicrobial resistance were also investigated. L. monocytogenes, L. innocua, and L. ivanovii were detected in C. caretta, whilst only L. monocytogenes and L. innocua in C. mydas. Notable among the results is the lack of significant differences in gene distribution between human and sea turtle strains. Furthermore, potentially pathogenic strains of L. monocytogenes were found in sea turtles.
Full article
(This article belongs to the Special Issue Microorganisms and Diseases Associated with Aquatic Animals 2.0)
►▼
Show Figures
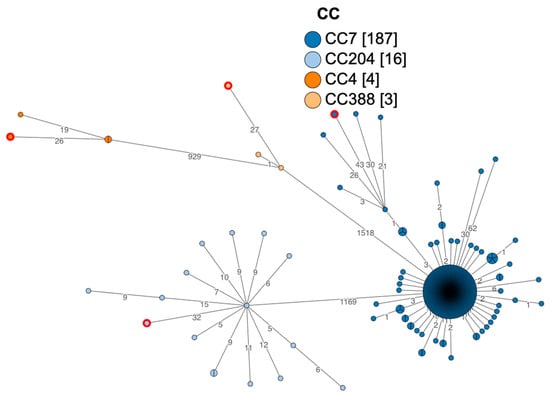
Figure 1
Open AccessArticle
Molecular and Serological Detection of Vector-Borne Pathogens Responsible for Equine Piroplasmosis in Europe between 2008 and 2021
by
Carla Wiebke Axt, Andrea Springer, Christina Strube, Clarissa Jung, Torsten J. Naucke, Elisabeth Müller and Ingo Schäfer
Microorganisms 2024, 12(4), 816; https://doi.org/10.3390/microorganisms12040816 - 17 Apr 2024
Abstract
Equine piroplasmosis (EP) is caused by Theileria (T.) equi and/or Babesia (B.) caballi. The aim was to assess the percentage of positive test results for EP in horses in Europe and to identify risk factors for pathogen contact/infection.
[...] Read more.
Equine piroplasmosis (EP) is caused by Theileria (T.) equi and/or Babesia (B.) caballi. The aim was to assess the percentage of positive test results for EP in horses in Europe and to identify risk factors for pathogen contact/infection. This study included results from PCR and competitive enzyme-linked immunosorbent assay testing requested by European veterinarians between 2008 and 2021. Binary bivariate logistic regression was used to analyze risk factors. A total of 4060 horses were included. PCR testing was positive in 9.7% (154/1589), serology for T. equi in 15.2% (393/2591) and for B. caballi in 6.8% (175/2578). The odds of positive serology increased by 6.8% (B. caballi, p = 0.008) and 9.5% (T. equi, p < 0.001) each year. Regionality had a statistically significant impact on PCR (Eastern p = 0.047/OR = 1.605; Southern p = 0.029/OR = 1.451; Central p = 0.007/OR = 0.617) and serological testing for T. equi (Southern p < 0.001/OR = 2.521; Central p < 0.001/OR = 0.537; Northern p = 0.003/OR = 0.462), as well as breeds on seroprevalence of B. caballi (heavy horses: p = 0.016/OR = 2.239) and T. equi (ponies: p = 0.007/OR = 0.340; warmbloods: p = 0.025/OR = 1.602). In conclusion, there was a significant geographical impact on the results of PCR and serology, consistent with known vector habitats. The rising numbers of horses tested serologically positive highlights the importance of surveillance.
Full article
(This article belongs to the Special Issue The Global Burden of Parasitic Diseases: Prevalence and Epidemiology)
►▼
Show Figures
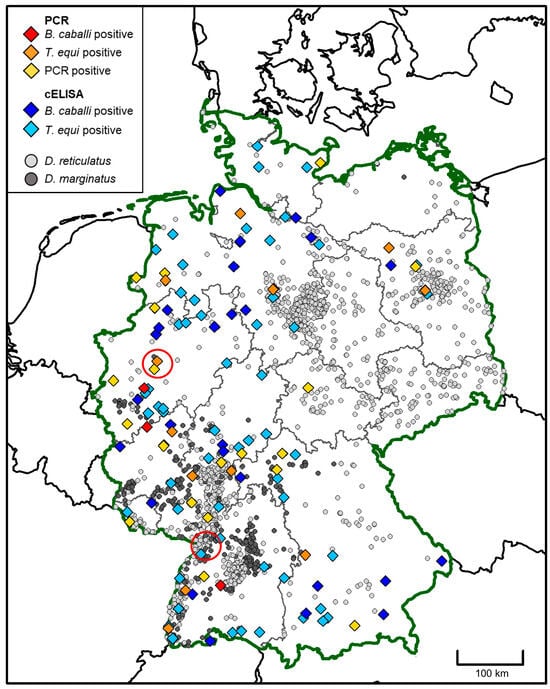
Figure 1

Journal Menu
► ▼ Journal Menu-
- Microorganisms Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biology, Ecologies, Forests, Microorganisms, Plants
Litter Decompositions: From Individuals to Ecosystems
Topic Editors: Wen Zhou, Guihua LiuDeadline: 30 April 2024
Topic in
Aquaculture Journal, Fishes, Microorganisms, Water
Women in Aquaculture Research
Topic Editors: Camino Ordás, Patrícia Díaz-RosalesDeadline: 30 June 2024
Topic in
Agronomy, Environments, Microorganisms, Pollutants, Sustainability, Water
Soil and Water Pollution Process and Remediation Technologies, 2nd Volume
Topic Editors: Hongbiao Cui, Ru Wang, Yu Shi, Haiying Lu, Lin ChenDeadline: 15 July 2024
Topic in
Antibiotics, Antioxidants, JoF, Microbiology Research, Microorganisms
Redox in Microorganisms, 2nd Edition
Topic Editors: Michal Letek, Volker BehrendsDeadline: 31 July 2024

Conferences
Special Issues
Special Issue in
Microorganisms
Plant Growth—Promoting Bacteria and Plant–Soil Interactions in Harsh Environments, 2nd Edition
Guest Editors: Blanca R. López, Luz De-BashanDeadline: 25 April 2024
Special Issue in
Microorganisms
Plant and Human Probiotics: Consequences on the Autochthonous Microbiota
Guest Editors: Elisa Gamalero, Elisa Bona, Giorgia Novello, Francesco VuoloDeadline: 30 April 2024
Special Issue in
Microorganisms
Human Skin Microbiota 2.0
Guest Editors: Holger Brüggemann, Rolf LoodDeadline: 15 May 2024
Special Issue in
Microorganisms
The Urban Microbiome
Guest Editors: Haruo Suzuki, Soojin JangDeadline: 31 May 2024
Topical Collections
Topical Collection in
Microorganisms
Trends in Yeast Biochemistry and Biotechnology
Collection Editors: Seiji Shibasaki, Mitsuyoshi Ueda
Topical Collection in
Microorganisms
Epidemiology and Pathogenicity of Animal-Adapted Streptococci
Collection Editors: Peter Valentin-Weigand, Marcus Fulde
Topical Collection in
Microorganisms
Biodegradation and Environmental Microbiomes
Collection Editors: Shuangjiang Liu, Hongzhi Tang, Jiandong Jiang, Xiaolei Wu